Tutorials
ECAL 2015 provides a venue for a number of tutorials, which are run and administered independently from the main conference. If you are interested in attending one of the tutorials, please visit the relevant website below. For tutorials that do not currently have a website, please contact the tutorial organiser for more details.
|
Physarum Machines: Slime Mould Computing and Novel Computing Substrates Tuesday 21st July, 10:00 - 13:00 Plasmodium of acellular slime mould Physarum polycephalum is a gigantic single cell visible by unaided eye. The cell shows a rich spectrum of behavioural morphological patterns in response to changing environmental conditions. Given data represented by chemical or physical stimuli we can employ and modify the behaviour of the slime mould to make it solve a range of computing and sensing tasks. We invited contributions on software, hardware and wetware realisations of slime mould computers. Exemplar topics of the submissions could be as follows: Hybrid wetware+hardware devices, Slime mould based electronic, Physarum inspired algorithms on graphs and networks, Computing models and prototypes of slime mould computers, Physarum inspired calculi and logic, Slime mould inspired algorithms of evacuation, Computing oriented biology of slime mould, Neuromorphic devices made of the slime mould,Cognitive functions modelled with the slime mould, Physarum in arts and music, Slime mould sensors, Cellular automata and particle based models of Physarum, Physarum inspired robots, Physarum controllers, Novel materials inspired by slime mould, Physarum and transport networks, Slime mould intelligence, Space exploration with slime mould. Run in conjunction with a workshop and a special session of the main conference. |
Organised by: |
|
Introduction to Complex Networks Modeling and Analysis with Python/NetworkX Monday 20th July, 16:00 - 18:00 The science of complex networks has been increasing its relevance to a wide variety of scientific disciplines over the last decade. It offers concepts and tools to help us understand various complex systems in nature from a viewpoint of connection and interaction. Subjects being studied range from the way cells function, to the ways neurons connect and configure brains, to the ways societies and economies grow and behave. This tutorial will provide a quick introduction to Python and NetworkX, a very powerful computational toolkit for modeling, analyzing and simulating complex networks. Specific topics to be introduced will include: Data structure and manipulation in Python/NetworkX, data import and export, network visualization, network analysis, generative network models, and dynamic simulations of complex networks. Instructions will be given through many hands-on activities, so participants should bring their own laptops. Prior knowledge of Python is helpful. |
Organised by: |
|
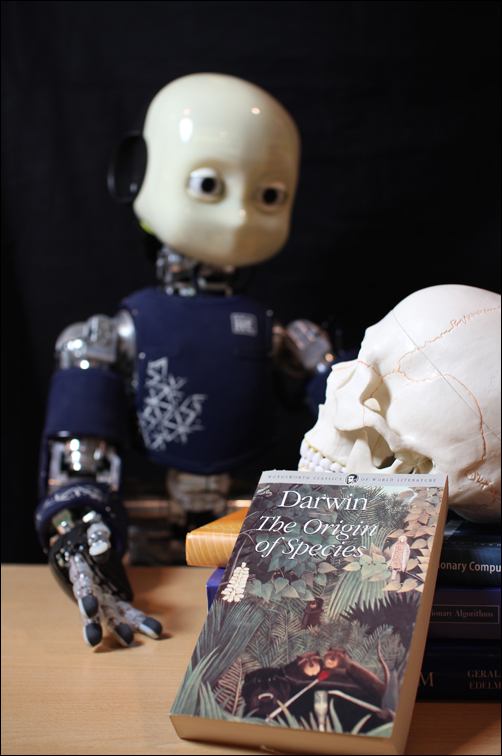
Aevol Tutorial Thursday 23rd July, 10:00 - 13:00 Aevol is a digital genetics model in which populations of digital organisms with a realistic genomic structure undergo variation and selection, creating a Darwinian dynamics. In this tutorial we will provide potential users with all the necessary information to test the software, as well as design and start their own experiments. In the first part of the tutorial, we will present the structure of the software and the artificial chemistry used to compute a phenotype from a genotype. We will also insist on the "in silico experimental evolution" methodology and on the similarities with in vivo experimental evolution. The second part will be a mini-lab during which the participants will have a chance to run either some classical evolutionary experiments or new ones of their own design. Finally, we will present the general overview of Aevol's code structure in order to help more advanced users to develop extensions of the initial model. |
Organised by:
|
|
Introduction to JIDT: An information-theoretic toolkit for studying the dynamics of complex systems Monday 20th July, 13:30 - 15:30 Complex systems in Artificial Life are increasingly being viewed as distributed information processing systems, resulting in a strong uptake of information-theoretic measures for analysis of dynamics in these fields. This tutorial will review the use of these measures, and then introduce participants to an open-source toolkit (JIDT) for conducting such analysis. Principally, JIDT implements the transfer entropy, (conditional) mutual information and active information storage, for both discrete and continuous-valued data. The toolkit can be used directly in Java, Matlab, Octave, Python, etc. Participants will learn how to install JIDT, how to get started with typical usage scenarios, and where to seek further support information. Ability to run code in your preferred environment (Java, Matlab/Octave, Python) is assumed, and to fully participate users should bring their own laptop (with that environment installed). |
Organised by: |
|
Evolutionary Robotics Thursday 23rd July, 14:00 - 17:25 In the same way as evolution shapes animals, is it possible to use artificial evolution to automatically design robots? This attractive vision gave birth to Evolutionary Robotics (ER), an interdisciplinary field that incorporates ideas from Biology, Robotics and Artificial Intelligence. Within the last 20 years, Evolutionary Robotics explored many research avenues, from understanding Natural Evolution thanks to ER models, to co-designing robots' bodies and brains. This tutorial will give an overview of the various questions addressed in ER, relying on examples from the literature. Past achievements and major contributions, as well as specific challenges in ER will be described. The tutorial will in particular focus on: what is evolutionary robotics?; fitness function design and the influence of selection pressure; encodings of controllers and morphologies; evolution for physical robots and the reality gap; embodied evolution and collective robotics systems; open questions and future challenges. |
Organised by:
|
|
An Introduction to the use of Proper Statistical Analysis for ALife Experiments Wednesday 22nd July, 10:00 - 13:00 It is becoming widely accepted that experimental results without proper statistical analysis must be considered anecdotal, and may even be wholly inaccurate. Increasingly, if your paper has inadequate statistical analysis, it will be rejected. This introductory tutorial aims at refreshing your memory on the basic statistical techniques used when analyzing experiments. Introductory topics are covered, starting with the T-test, confidence intervals, and non-parametric statistics, and progressing to statistical model building through regression (linear and polynomial), and Analysis of Variance (ANOVA). All ideas and techniques are intuitively presented using Excel and R examples and through the extensive use of graphics and animation … concentrating on concepts, not equations. |
Organised by: |